I'm currently a research scientist at Adobe. My research interests include physically-based animation and physical simulation. I aim to make it easier and more efficient to simulate various phenomena, and also to make it easier to create physically realistic animations.
Internships @ Adobe Research: If you are a PhD student interested in doing a research internship at Adobe and/or collaborating on a project with me, send an e-mail. It will be helpful if you include a CV and a summary of your current research interests.
-
Ph.D., Computer Science
Cornell University
2011-2016 -
B.S., Computer Engineering
University of Massachusetts Amherst
2005-2009
- SIGGRAPH Technical Papers committee member: 2024, 2025
- Technical paper reviewer: SIGGRAPH, SIGGRAPH Asia, ACM TON, ECCV, TVCG, IEEE VR, CGF, Pacific Graphics
- Member of Adobe Employee Community Fund grant committee
-
Cornell CS Student Brown
Bag Czar
2013-2015
-
Volunteer with
Expand Your Horizons
Helped organize an educational workshop for middle school students
Spring 2012
-
Senior Research Scientist at Adobe Research
2018-Present -
Research Scientist at Adobe Research
2016-2018 -
Research Intern at Disney Research Boston
2015 (summer) -
Software Engineer in the MIT Lincoln Laboratory Weather Sensing Group
Developed weather prediction algorithms, distributed real-time systems
2009-2011 -
Software Engineering Intern at Raytheon
Summer 2008 -
Software Engineering Intern at DEKA Research and Development
Embedded systems development on several medical devices
Summers and winters 2006-2008
One of the main projects running some of my code is the DEKA Arm
Videos of coverage from 60 Minutes and IEEE Spectrum
- Dynamic Animation in Adobe Express
- Spatial Audio in Aero
- Magnets in Character Animator
- Colliding Particles in Character Animator
- Rigid Body Physics in Character Animator
- Ambisonics in Premiere Pro
journal = {Computer Graphics Forum},
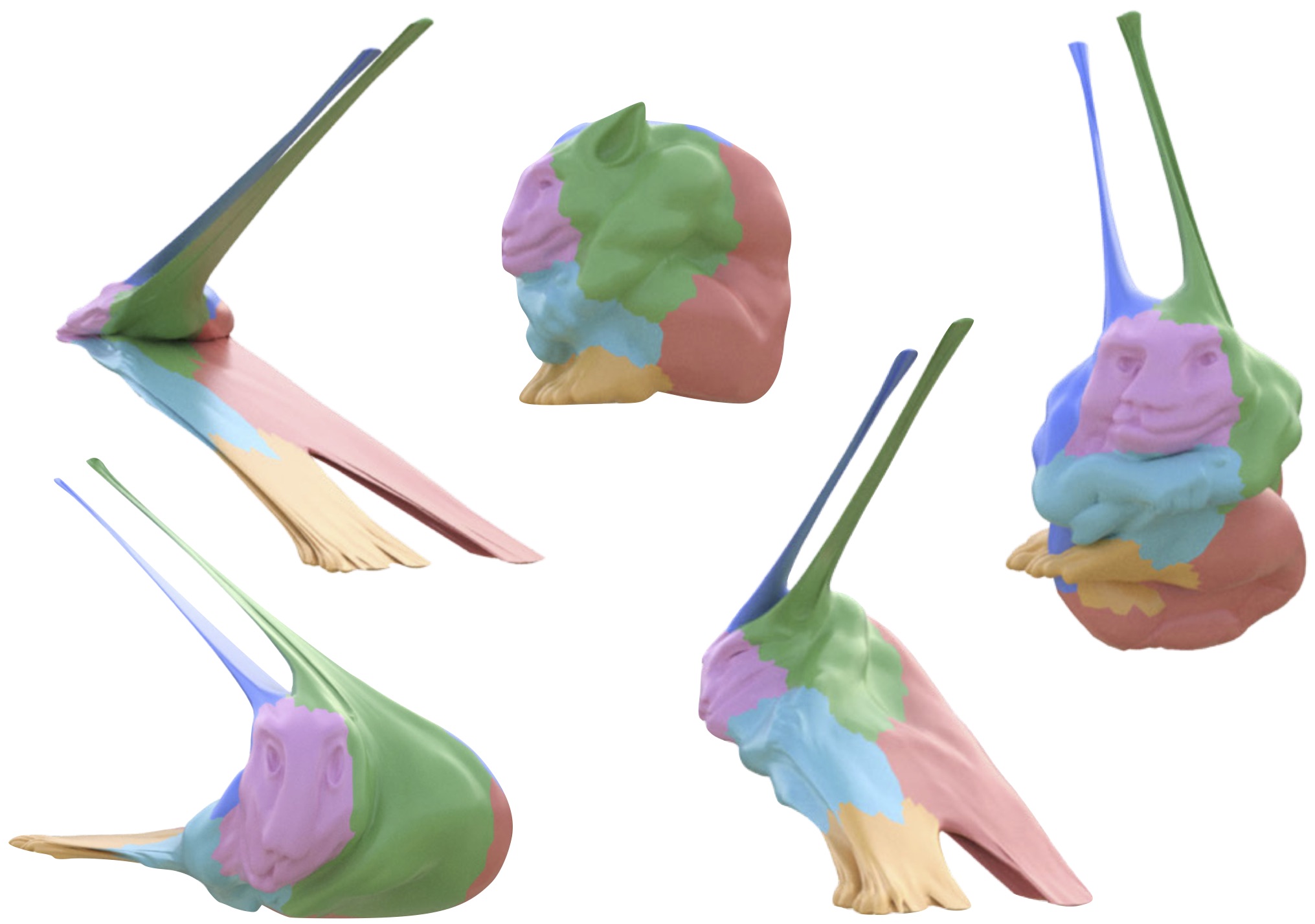
title = {{How to Train Your Dragon: Automatic Diffusion-Based Rigging for Characters with Diverse Topologies}},
author = {Gu, Zeqi and Liu, Difan and Langlois, Timothy and Fisher, Matthew and Davis, Abe},
year = {2025},
publisher = {The Eurographics Association and John Wiley & Sons Ltd.},
ISSN = {1467-8659},
DOI = {10.1111/cgf.70016}
}
author = {Liu, Vivian and Kazi, Rubaiat Habib and Wei, Li-Yi and Fisher, Matthew and Langlois, Timothy and Walker, Seth and Chilton, Lydia},
title = {LogoMotion: Visually-Grounded Code Synthesis for Creating and Editing Animation},
year = {2025},
isbn = {9798400713941},
publisher = {Association for Computing Machinery},
address = {New York, NY, USA},
url = {https://doi.org/10.1145/3706598.3714155},
doi = {10.1145/3706598.3714155},
booktitle = {Proceedings of the 2025 CHI Conference on Human Factors in Computing Systems},
articleno = {157},
numpages = {16},
series = {CHI '25}
}
author = {Xue, Kangrui and Wang, Jui-Hsien and Langlois, Timothy and James, Doug},
title = {WaveBlender: Practical Sound-Source Animation in Blended Domains},
year = {2024},
isbn = {9798400711312},
publisher = {Association for Computing Machinery},
address = {New York, NY, USA},
url = {https://doi.org/10.1145/3680528.3687696},
doi = {10.1145/3680528.3687696},
booktitle = {SIGGRAPH Asia 2024 Conference Papers},
articleno = {134},
numpages = {10},
series = {SA '24}
}
author = {Chen, Yixin and Levin, David and Langlois, Timothy},
title = {Fluid Control with Laplacian Eigenfunctions},
year = {2024},
url = {https://doi.org/10.1145/3641519.3657468},
doi = {10.1145/3641519.3657468},
booktitle = {ACM SIGGRAPH 2024 Conference Papers},
articleno = {44},
numpages = {11},
location = {Denver, CO, USA},
series = {SIGGRAPH '24}
}
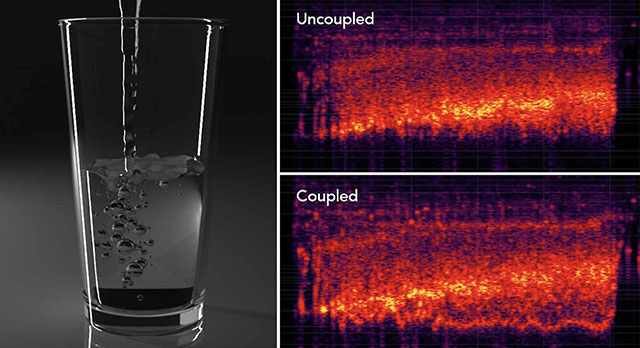
author = {Xue, Kangrui and Aronson, Ryan M and Wang, Jui-Hsien and Langlois, Timothy R and James, Doug L},
title = {Improved Water Sound Synthesis using Coupled Bubbles},
year = {2023},
issue_date = {August 2023},
publisher = {Association for Computing Machinery},
address = {New York, NY, USA},
volume = {42},
number = {4},
journal = {ACM Trans. Graph.},
month = {aug},
}
author = {Cui, Qiaodong and Langlois, Timothy and Sen, Pradeep and Kim, Theodore},
title = {Spiral-Spectral Fluid Simulation},
year = {2021},
issue_date = {December 2021},
publisher = {Association for Computing Machinery},
address = {New York, NY, USA},
volume = {40},
number = {6},
journal = {ACM Trans. Graph.},
month = {dec},
articleno = {202},
numpages = {16}
}
Our algorithm explicitly models the curved trajectories traced by rigid bodies in both collision detection and response. For collision detection, we propose a conservative narrow phase collision detection algorithm for curved trajectories, which reduces the problem to a sequence of linear CCD queries with minimal separation. For time integration and contact response, we extend the recently proposed incremental potential contact framework to reduced coordinates and rigid body dynamics.
We introduce a benchmark for rigid body simulation and show that our approach, while less efficient than alternatives, can robustly handle a wide array of complex scenes, which cannot be simulated with competing methods, without requiring per-scene parameter tuning.
title = {Intersection-free Rigid Body Dynamics},
author = {Zachary Ferguson and Minchen Li and Teseo Schneider and Francisca Gil-Ureta and Timothy Langlois and Chenfanfu Jiang and Denis Zorin and Danny M. Kaufman and Daniele Panozzo},
year = 2021,
journal = {ACM Transactions on Graphics (SIGGRAPH)},
volume = 40,
number = 4,
articleno = 183
}
Constructed with a custom nonlinear solver, IPC enables efficient res- olution of time-stepping problems with separate, user-exposed accuracy tolerances that allow independent specification of the physical accuracy of the dynamics and the geometric accuracy of surface-to-surface conformation. This enables users to decouple, as needed per application, desired accuracies for a simulation’s dynamics and geometry.
The resulting time stepper solves contact problems that are intersection- free (and thus robust), inversion-free, efficient (at speeds comparable to or faster than available methods that lack both convergence and feasibility), and accurate (solved to user-specified accuracies). To our knowledge this is the first implicit time-stepping method, across both the engineering and graphics literature that can consistently enforce these guarantees as we vary simulation parameters.
In an extensive comparison of available simulation methods, research libraries and commercial codes we confirm that available engineering and computer graphics methods, while each succeeding admirably in custom- tuned regimes, often fail with instabilities, egregious constraint violations and/or inaccurate and implausible solutions, as we vary input materials, contact numbers and time step. We also exercise IPC across a wide range of existing and new benchmark tests and demonstrate its accurate solution over a broad sweep of reasonable time-step sizes and beyond (up to h = 2s) across challenging large-deformation, large-contact stress-test scenarios with meshes composed of up to 2.3M tetrahedra and processing up to 498K contacts per time step. For applications requiring high-accuracy we demon- strate tight convergence on all measures. While, for applications requiring lower accuracies, e.g. animation, we confirm IPC can ensure feasibility and plausibility even when specified tolerances are lowered for efficiency.
author = {Minchen Li and Zachary Ferguson and Teseo Schneider and Timothy Langlois and Denis Zorin and Daniele Panozzo and Chenfanfu Jiang and Danny M. Kaufman},
title = {Incremental Potential Contact: Intersection- and Inversion-free Large Deformation Dynamics},
journal = {ACM Transactions on Graphics},
year = {2020},
volume = {39},
number = {4} }
author={Tang, Zhenyu and Bryan, Nicholas J and Li, Dingzeyu and Langlois, Timothy R and Manocha, Dinesh},
journal={IEEE Transactions on Visualization and Computer Graphics},
title={Scene-Aware Audio Rendering via Deep Acoustic Analysis},
year={2020},
volume={26},
number={5},
pages={1991-2001} }
title = {Decomposed Optimization Time Integrator for Large-Step Elastodynamics},
author = {Li, Minchen and Gao, Ming and Langlois, Timothy R. and Jiang, Chenfanfu and Kaufman, Danny M.},
journal = {ACM Transactions on Graphics},
volume = {38},
number = {4},
year = {2019} }
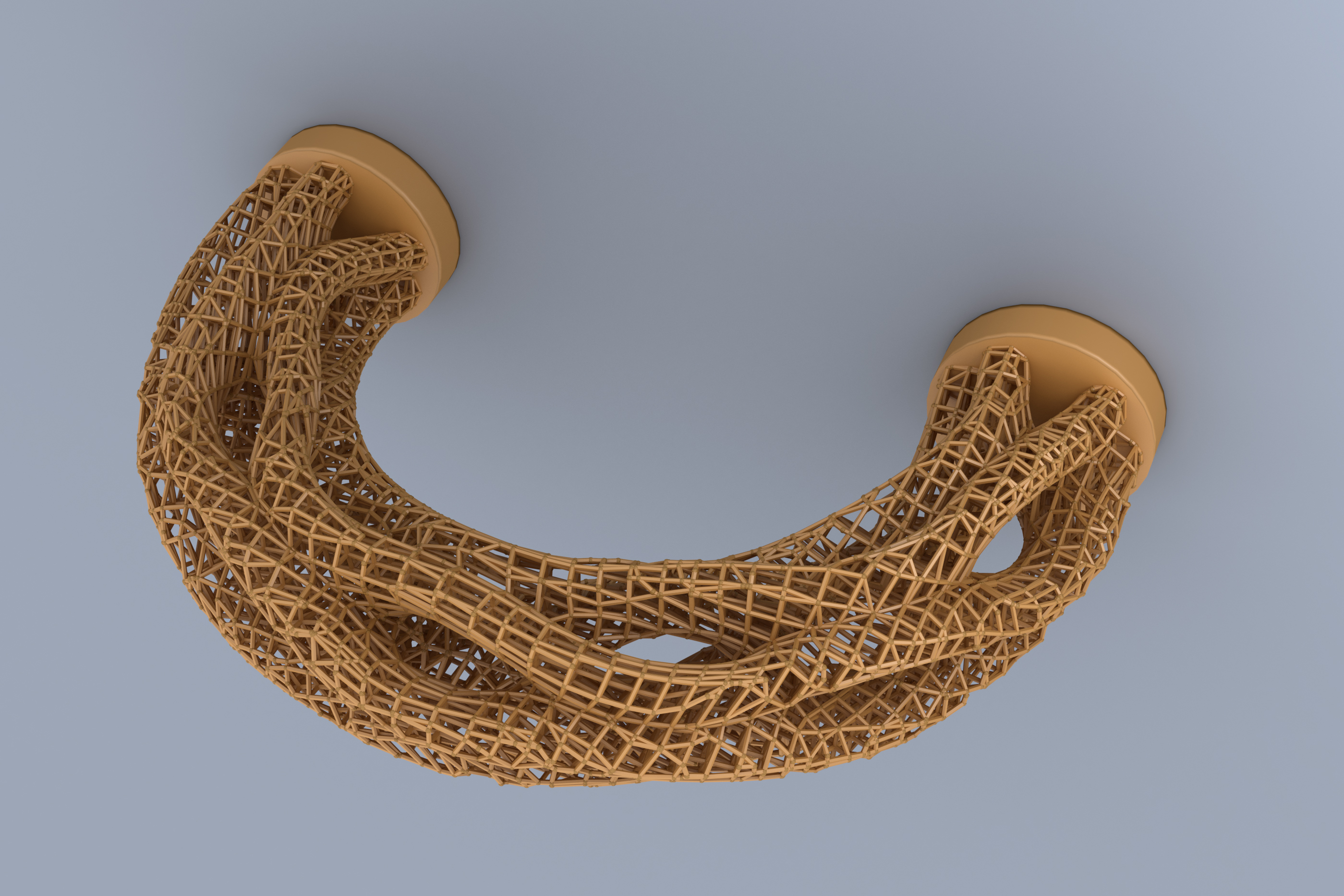
author = {Arora, Rahul and Jacobson, Alec and Langlois, Timothy R. and Huang, Yijiang and Mueller, Caitlin and Matusik, Wojciech and Shamir, Ariel and Singh, Karan and Levin, David I.W.},
title = {Volumetric Michell Trusses for Parametric Design \& Fabrication},
booktitle = {Proceedings of the 3rd ACM Symposium on Computation Fabrication},
series = {SCF '19},
year = {2019},
location = {Pittsburgh, PA, USA},
numpages = {12},
publisher = {ACM},
address = {New York, NY, USA} }
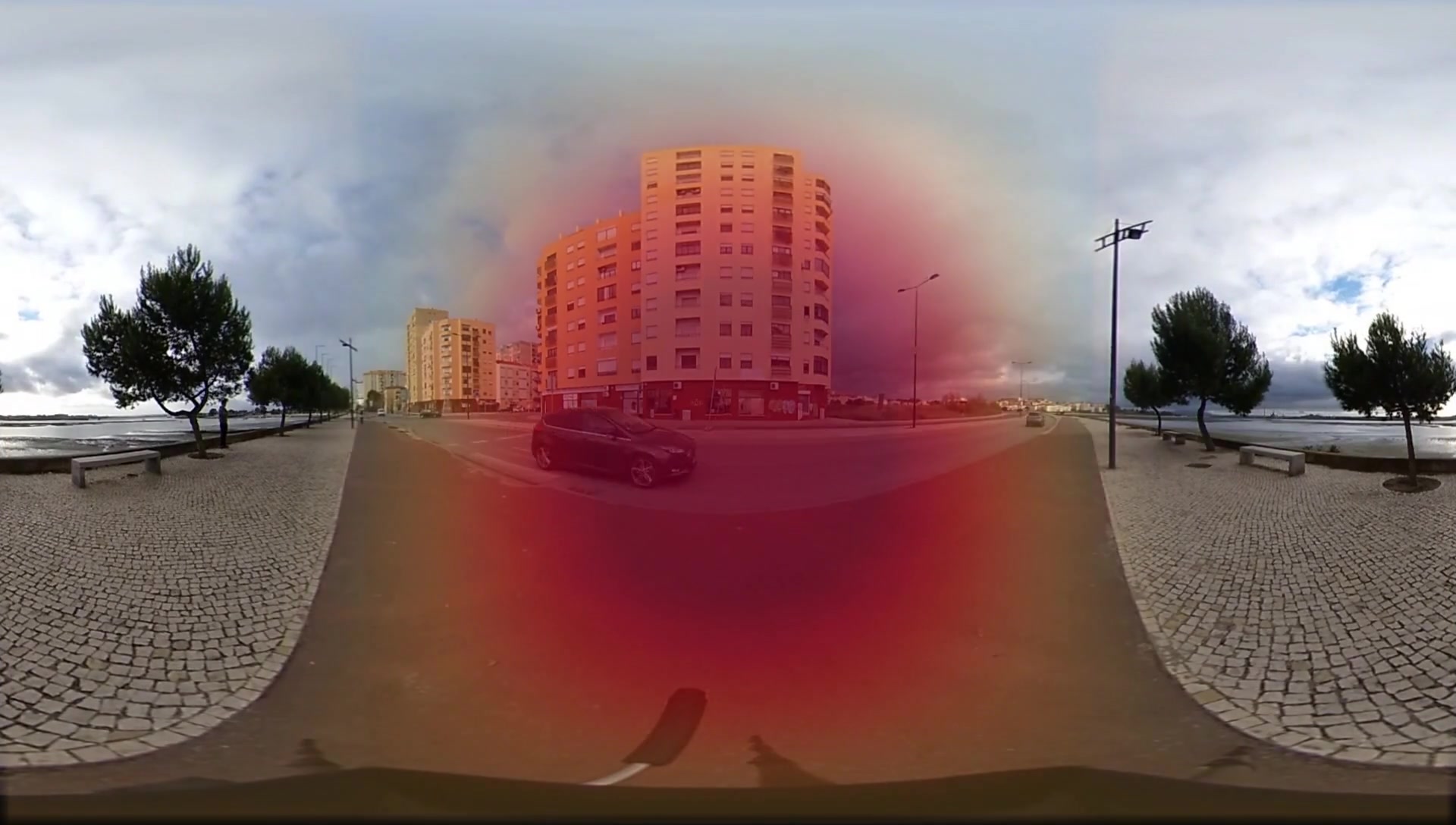
title = {Self-Supervised Generation of Spatial Audio for 360\deg Video},
author = {Pedro Morgado, Nuno Vasconcelos, Timothy Langlois and Oliver Wang},
booktitle = {Neural Information Processing Systems (NIPS)},
year = {2018} }
title = {Toward Wave-based Sound Synthesis for Computer Animation},
author = {Wang, Jui-Hsien and Qu, Ante and Langlois, Timothy R. and James, Doug L.},
journal = {ACM Trans. Graph.},
volume = {37},
number = {4},
year = {2018}, }
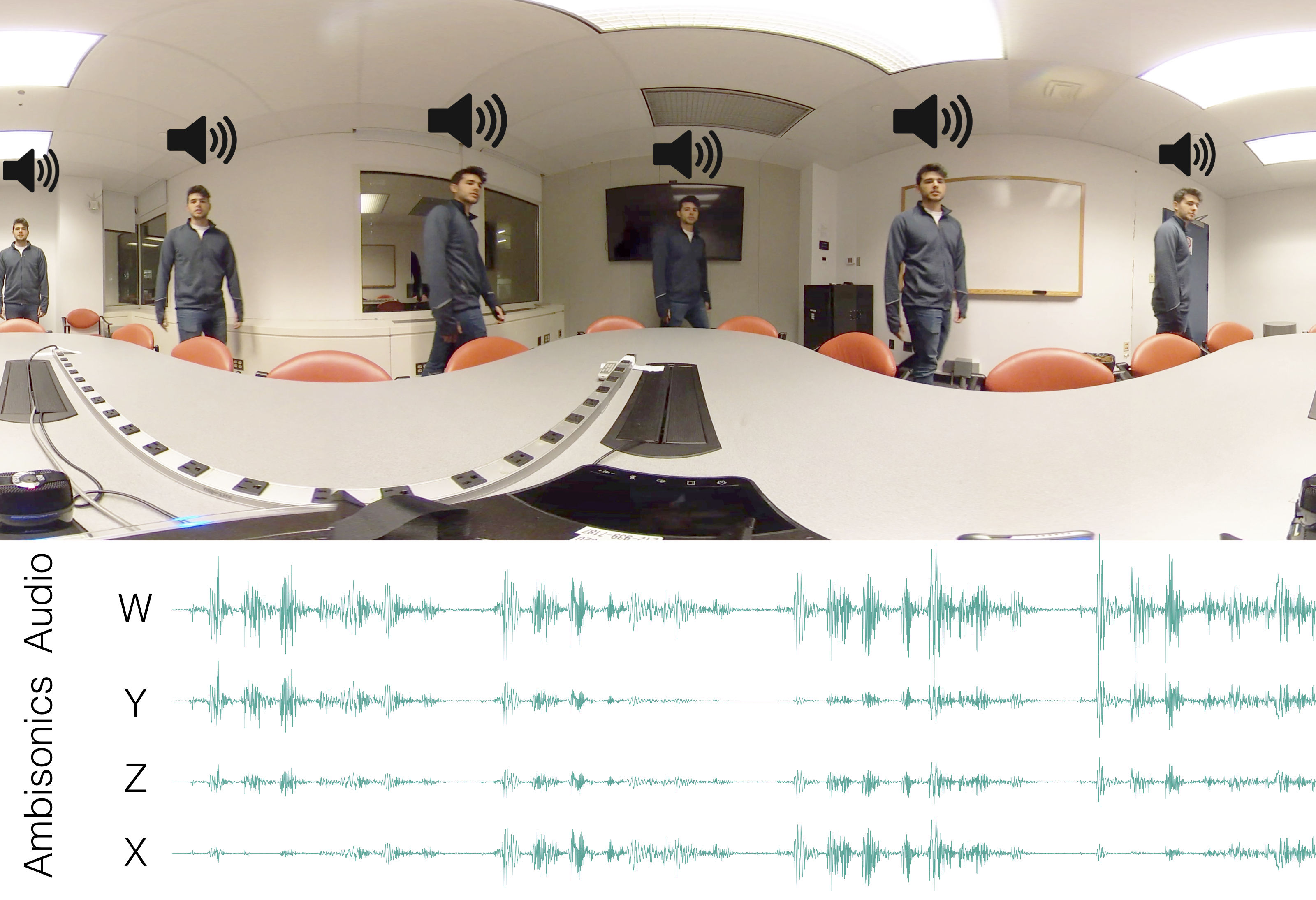
title = {Scene-Aware Audio for 360\textdegree{} Videos},
author = {Li, Dingzeyu and Langlois, Timothy R. and Zheng, Changxi},
journal = {ACM Trans. Graph.},
volume = {37},
number = {4},
year = {2018}, }
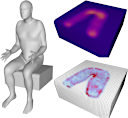
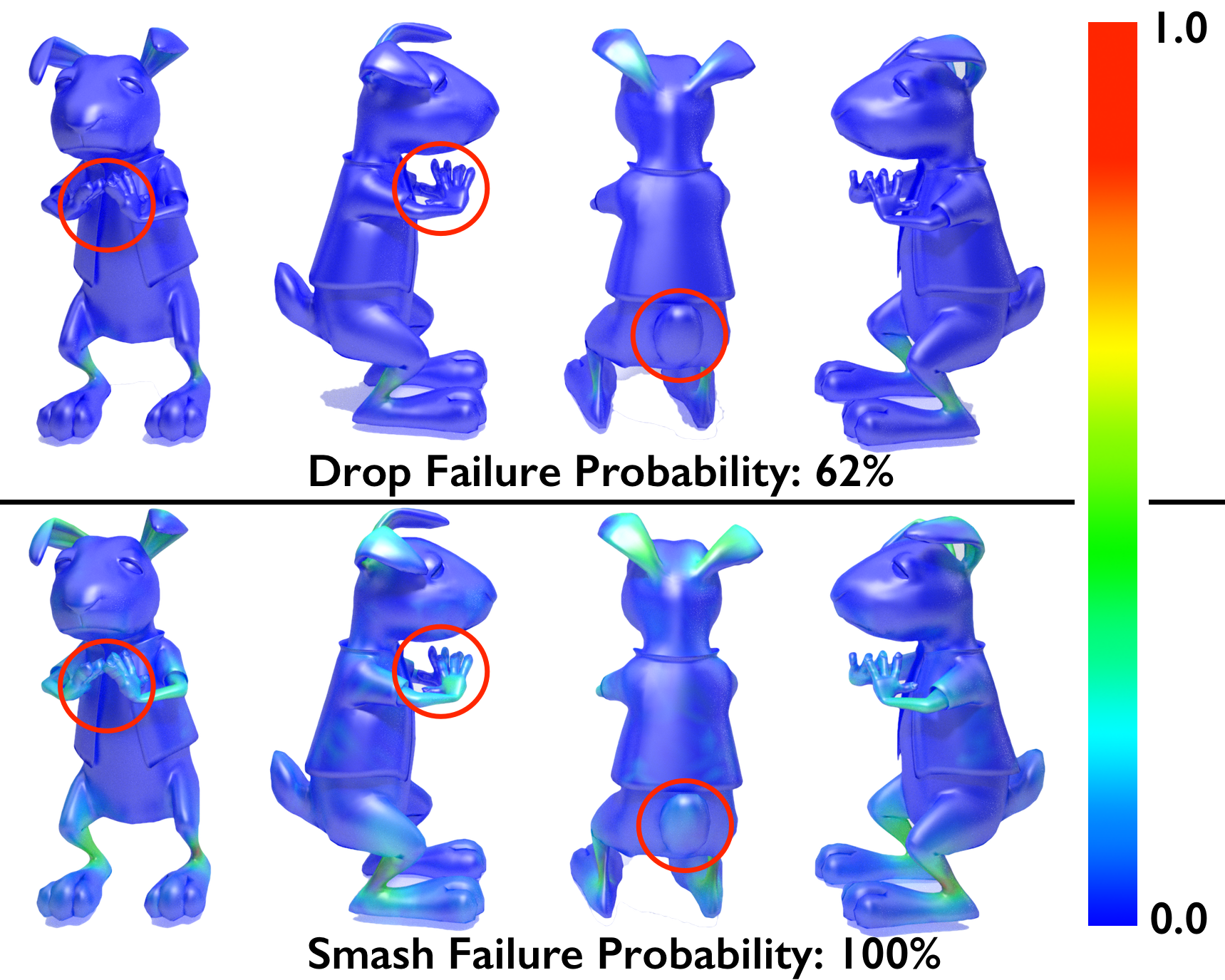
author = {Langlois, Timothy and Shamir, Ariel and Dror, Daniel and Matusik, Wojciech and Levin, David I. W.},
title = {Stochastic Structural Analysis for Context-aware Design and Fabrication},
journal = {ACM Trans. Graph.},
issue_date = {November 2016},
volume = {35},
number = {6},
month = nov,
year = {2016},
issn = {0730-0301},
pages = {226:1--226:13},
articleno = {226},
numpages = {13},
url = {http://doi.acm.org/10.1145/2980179.2982436},
doi = {10.1145/2980179.2982436},
acmid = {2982436},
publisher = {ACM},
address = {New York, NY, USA},
keywords = {FEM, computational design, structural analysis}, }
author = {Timothy R. Langlois and Changxi Zheng and Doug L. James},
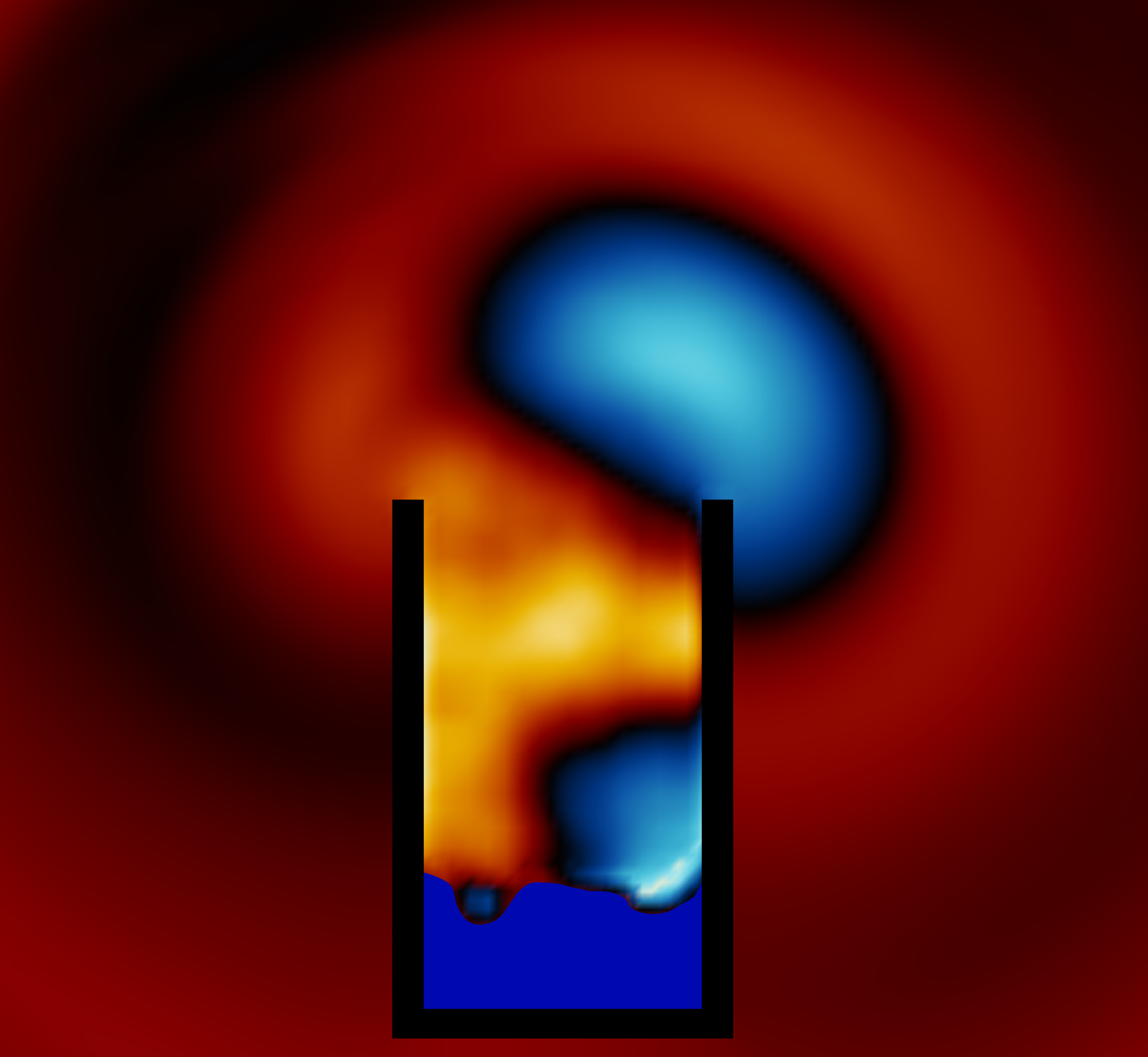
title = {Toward Animating Water with Complex Acoustic Bubbles},
journal = {ACM Transactions on Graphics (Proceedings of SIGGRAPH 2016)},
year = {2016},
volume = {35},
number = {4},
month = Jul,
doi = {10.1145/2897824.2925904}
url = {http://www.cs.cornell.edu/projects/Sound/bubbles}
}
author = {Timothy R. Langlois and Steven S. An and Kelvin K. Jin and Doug L. James},
title = {Eigenmode Compression for Modal Sound Models},
journal = {ACM Transactions on Graphics (Proceedings of SIGGRAPH 2014)},
year = {2014},
volume = {33},
number = {4},
month = Aug,
doi = {10.1145/2601097.2601177}
url = {http://www.cs.cornell.edu/projects/Sound/modec}
}
author = {Timothy R. Langlois and Doug L. James},
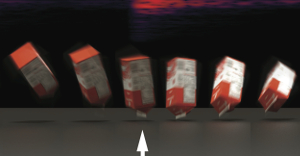
title = {Inverse-Foley Animation: Synchronizing rigid-body motions to sound},
journal = {ACM Transactions on Graphics (Proceedings of SIGGRAPH 2014)},
year = {2014},
volume = {33},
number = {4},
month = Aug,
doi = {10.1145/2601097.2601178}
url = {http://www.cs.cornell.edu/projects/Sound/ifa}
}
year={2010},
isbn={978-1-4419-5912-6},
booktitle={Advances in Computational Biology},
volume={680},
series={Advances in Experimental Medicine and Biology},
editor={Arabnia, Hamid R.},
doi={10.1007/978-1-4419-5913-3_39},
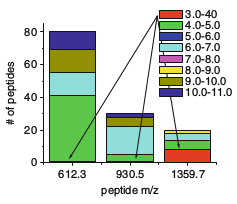
title={Protein Identification Using Receptor Arrays and Mass Spectrometry},
url={http://dx.doi.org/10.1007/978-1-4419-5913-3_39},
publisher={Springer New York},
keywords={Receptor; Array; Mass; Spectrometry; Protein; Identification; Isoelectric; Point},
author={Langlois, Timothy R. and Vachet, Richard W. and Mettu, Ramgopal R.},
pages={343-351},
language={English}
}
Pool Table Analyzer
My group's Senior Design Project at UMass. We designed and built a system which watched a game of pool using webcams, suggested the best shot to the player, and assisted them with aiming the cue stick, all in realtime. More details here.